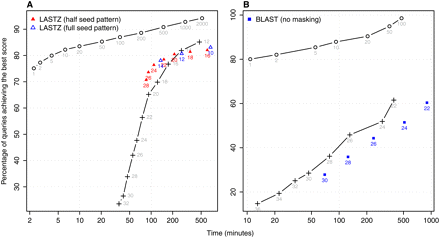
LASTZ and BLASTN use fixed-length seeds and present similar performance to LAST with fixed-length seeds. (Circles) LAST adaptive seeds; (crosses) LAST fixed-length seeds. (A) P. yoelii contigs are aligned to P. falciparum chromosomes using the same score matrix as in Figure 3C. Triangles show performance of LASTZ executed with corresponding parameters for different LASTZ-seed lengths. (B) A simpler match/mismatch scoring scheme is used. Squares present the performance of BLASTN and the numbers correspond to BLASTN-seed lengths.











