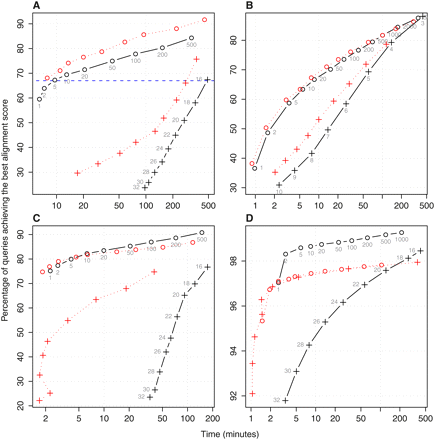
Performance comparison of adaptive seeds (circles) versus fixed-length seeds (crosses). Black denotes results obtained for contiguous seeds and unmasked sequences (l or f parameters are shown next to each data point). Red shows the effect of spaced seeds, subset seeds, or repeat-masking for the sequence alignment of: (A) H. sapiens promoters to the M. musculus genome (and spaced seed 111010010100110); (B) D. melanogaster protein sequences to those of C. elegans (and subset seeds); (C) P. yoelii contigs to the P. falciparum genome (and soft-masking with Tandem Repeats Finder); and (D) short DNA reads from 454 Life Sciences (Roche) GS 20 for A. thaliana against its genome (and soft-masking with WindowMasker).











