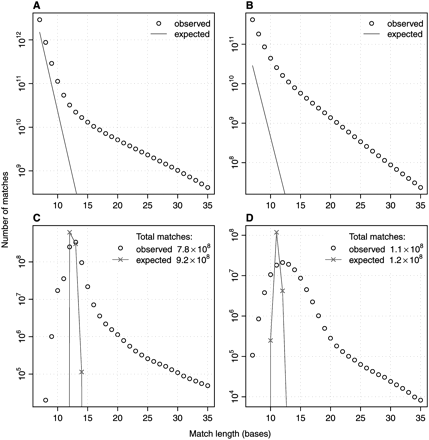
Number of exact matches between genomic sequences as a function of match length. (A) Matches of size 7–35 bases between the human X chromosome (151 million bases) and the mouse X chromosome (162 million bases). (B) Matches between the genomes of P. falciparum (23 million bases) and P. yoelii (20 million bases). (C) Matches between the human X chromosome and the mouse X chromosome, of seeds that occur at most 10 times in the mouse X chromosome. (D) Matches between P. falciparum and P. yoelii, of seeds that occur at most 10 times in P. falciparum. Lines show expected frequencies for uniformly random sequences.











