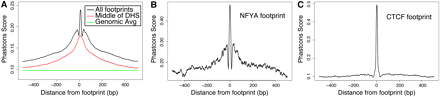
Conservation of sequence in and around DNase I footprints. (A) In general, footprints contain a strong sequence conservation signal with a nearby “shoulder” of conservation around all footprinted regions (black). Between the conservation peak and shoulder is a region with a marked decrease in conservation. This conservation pattern is not detected when the signal is centered on DNase I hypersensitive sites (red). The average conservation signal across the genome is shown in green. (B) The conservation pattern for a single factor, NFYA, displays the characteristic drop in conservation surrounding the footprint. (C) This conservation pattern is not detected around CTCF footprints, which shows relatively little conservation outside the highly conserved footprint. See also Supplemental Figure 6.











