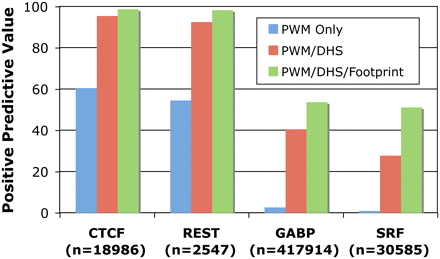
Accuracy of footprinting model. Positive predictive value (PPV) was calculated for predictions of four factors: CTCF, REST, GABP, and SRF. True-positives were determined by ChIP-seq peaks with a matching motif, while true-negatives were determined by motifs without corresponding ChIP-seq peaks. PPV is shown for predictions using only PWMs (all PWMs are considered an actual binding site), PWMs that map within DHS sites (all PWMs within a DNase I hypersensitive site are considered actual binding sites, while those PWMs outside of DNase I hypersensitive sites are considered negatives), and PWMs that map within footprints (all PWMs within a footprint are considered actual binding sites while those PWMs outside of footprints are considered negatives). The total number of PWMs mapped to the genome for each factor is listed in parentheses.











