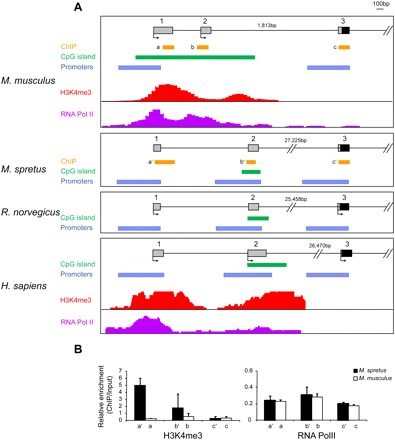
Comparison of Clcn4 5′ end between eutherian mammals. (A) Map locations of putative promoters (P1, P2, and P3) identified by Genomatix analysis, of CpG islands, and of known regions enriched in H3K4me3 and PolII, in M. musculus, M. spretus, rat, and human. Exon numbers (1, 2, 3) are at top. Chromatin enrichment data was downloaded to the UCSC browser for M. musculus (Gupta et al. 2010) and for human (ENCODE) (Celniker et al. 2009). Arrows indicate starts of transcription based on reported cDNAs (Flicek et al. 2010). (B) Chromatin analysis of Clcn4-2 in the Patski cell line using ChIP followed by quantitative PCR to determine enrichment relative to input using primers specific for M. musculus and M. spretus loci. Relative enrichment is shown for H3K4me3 and RNA PolII using primers at exons 1 (a/a′), 2 (b/b′), and 3 (c/c′) whose position is indicated in A (Supplemental Table S6).











