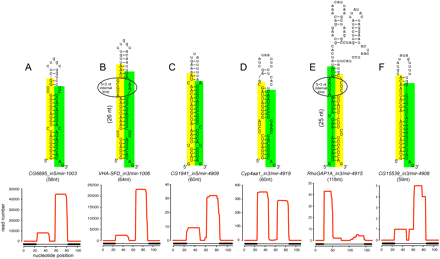
Examples of known and novel mirtrons in D. melanogaster. The abundant small RNAs derived from each hairpin are highlighted, green for the miRNA and yellow for the miRNA*. Below the secondary structures are plots that show the abundance of cloned small RNAs across the aggregate D. melanogaster small RNA data. The small RNA density is highest at either end of each intron, with typically one side accumulating to a higher level; often this is the 3′ arm, but occasionally it is the 5′ arm. The black boxes below the graph indicate the exon–intron boundaries. (A) CG6695_in5/mir-1003 is an example of a conserved, abundantly expressed mirtron with optimal features, including a straight short intronic hairpin with a 2-nt 3′ overhang. (B) Vha-SFD_in3/mir-1006 is an example of a conserved, abundant-expressed mirtron with a large asymmetric internal loop (5 + 2 nt). CG1941_in5 (C) and Cyp4aa1_in3 (D) are novel mirtrons with typical straight hairpins and compatible overhangs. (E) RhoGAP1A_in3 is an expressed mirtron with an unusually large, unstructured terminal loop, a large asymmetric internal loop (5 + 3 nt) and single nucleotide overhangs at its 5′ and 3′ ends. (F) CG15539_in3 exhibits convincing mirtron features, but is on the borderline of confident cloning evidence; nevertheless, its reads exhibit a characteristic 2-nt 3′ overhang on the Dicer-1-cleaved end.











