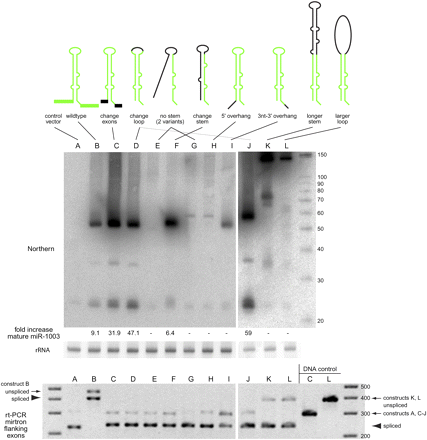
Structure-function analysis of mirtron biogenesis. (Top) S2 cells were transfected with UAS-mirtron and ub-Gal4 plasmids and RNA was isolated and subjected to Northern blot using an LNA probe antisense to miR-1003. Ethidium bromide staining of 5S rRNA is shown as a loading control. The fold increase in mature miR-1003 above control transfections is indicated below; (−) No substantial increase in miR-1003 level (>2 folds) was detected. (A) Control transfection using empty expression vector shows that S2 cells express a low level of the mirtron-derived miRNA miR-1003. (B) Introduction of mir-1003 expression plasmid, which includes portions of its endogenous flanking exons, yields strongly elevated pre-mir-1003 and mature miR-1003. Neither substitution of mir-1003 exonic context (C), nor replacement of its terminal loop (D), interferes with its biogenesis. Extensive mutation of its miRNA* arm abolishes production of miR-1003 (E,G), although a small amount of pre-miRNA is detected in the later case. However, extensive mutation while maintaining hairpin structure supports efficient mirtron biogenesis (F). (H) Introduction of a 5′ hairpin overhang abolishes small RNA production. (I) Extension of the 3′ hairpin overhang strongly impairs mirtron processing, although pre-miRNA accumulated. (J–L) Starting with a terminal loop mutant of mir-1003 (J, see also lane D), structured (K), and unstructured (L) hairpin extensions were introduced. Both constructs yielded substantial amounts of ∼150 nt pre-miRNA product, with higher levels of the fully duplexed intron (K); however, neither supported accumulation of mature miRNA. A ∼75-nt product corresponding to approximately half of the long hairpin intron accumulated; its biogenesis is not known. (Bottom) The same RNA samples used for Northern blotting at top were subjected to RT-PCR analysis to verify splicing accuracy of the mirtron variants. We observed weaker bands for the unspliced products and stronger bands for the spliced products; the DNA template controls at the right provide a size marker to gauge the unspliced amplification products. Note that the wild-type mir-1003 construct in its native CG6995 context includes more exon sequence than the other constructs, leading to the larger sizes of its RT-PCR products (“B”).











