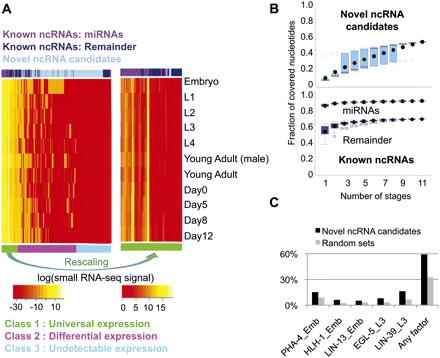
Expression patterns of the novel ncRNA candidates and binding signals of Pol II and 22 transcription factors around their genomic regions. (A) Expression patterns of known miRNA transcripts, other known ncRNA transcripts, and the candidate ncRNA bins based on our small RNA sequencing data at 11 developmental stages. Expression values are the log-transformed normalized read counts (DCPM, depth of coverage per million reads). Three subclasses formed according to the expression patterns are shown in the bottom row in three different colors. All known ncRNA transcripts are shown, while 1000 bins were randomly sampled from a total of 10,994 candidate ncRNA bins for this heat map visualization. The right panel shows a magnified view of class 1, with the colors rescaled to show the fluctuation of expression patterns across the different developmental stages. (B) Saturation plots of expressed known ncRNA transcripts and candidate ncRNA bin in different developmental stages. The fractions of expressed regions (with small RNA-seq signals stronger than the average signal of gold-standard intergenic regions) at the 11 developmental stages are computed using all possible combinations of the stages. The x-axis corresponds to the number of stages considered, and each point at a given number of stages corresponds to a different combination of stages. (C) Fractions of intergenic candidate ncRNA fragments potentially targeted by a selected subset of transcription factors. The total fractions targeted by any of the transcription factors in any of the stages are also shown. Each bar is labeled by the name of the transcription factor followed by the stage at which the binding experiment was performed. The bindings on random genome locations with the same size are also shown. EMB, embryo; YA, young adult.











