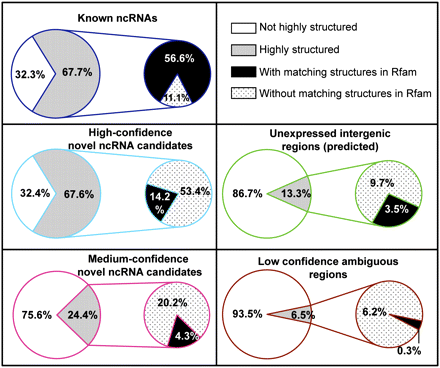
Structural properties of the known ncRNAs and novel ncRNA candidates. The percentages of highly structured ncRNA bins (with predicted Z-score of folding free energy less than the mean by at least one standard deviation) are shown for the known ncRNA bins and for high-confidence and medium-confidence candidate ncRNA bins. Among the highly structured bins, the percentages that overlap with structural homologs of Rfam families are also shown. The same calculations were performed for the bins predicted to be unexpressed intergenic regions and the low-confidence ambiguous regions (bins) for comparison.











