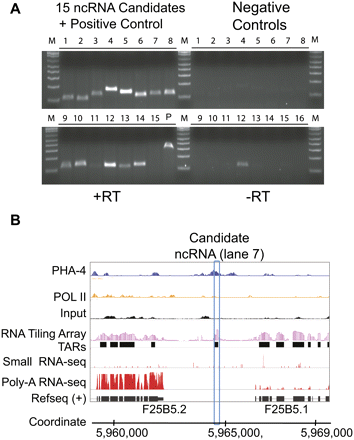
Validation of our novel ncRNA candidates. (A) Fifteen novel ncRNA candidates were tested using RT-PCR. Lanes 1–15 on the left (+RT) correspond to the PCR results of the novel ncRNA candidates, and lane 16 is the positive control, HEX-3. The right lanes (−RT) are the negative controls without reverse transcriptase. The PCR sizes are ∼150 bp in length. Fourteen of the candidates (except lane 15) were detected on the gel, among which 13 (except lane 11) showed a clear enrichment of expression signals in contrast to the negative control. (B) Example of a validated novel ncRNA candidate (ncRNA at lane 7 in A) with support from multiple information sources. The first and second rows (from top) correspond to the ChIP-seq reads from PHA-4 and Pol II, respectively. The heights of signals are normalized by their total mapped reads. The fourth and fifth rows are the log-transformed values of the tiling array in late embryo and its TARs. The bottom two rows are the reads from small RNA sequencing and the reads from poly-A+ RNA sequencing. The last row is the annotated genes from RefSeq.











