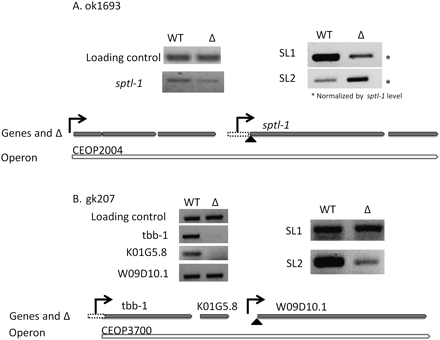
Effect of promoter deletions on the SL1/SL2 ratio. Diagrams at the bottom of each panel are to scale. They show pointed gray rectangles to indicate genes, hollow pointed empty rectangles to indicate operons and dotted boxes to indicate deletions. Black arrows indicate predicted locations of promoters and the triangle indicates the trans-splice site analyzed by RT-PCR. (A) RT-PCR in wild type (WT) and ok1693 (Δ). To make visualization of SL1/SL2 ratio easier, sptl-1 levels have been normalized to wild-type by adding 4× cDNA of the deletion strain based on the levels of mRNA for this gene (data not shown). (B) RT-PCR in wild type (WT) and gk207 (Δ). (Left) RT-PCR of mature mRNAs. (Right) RT-PCR of W09D10.1 trans-spliced to SL1 and SL2. The same level of cDNA was used for all PCRs.











