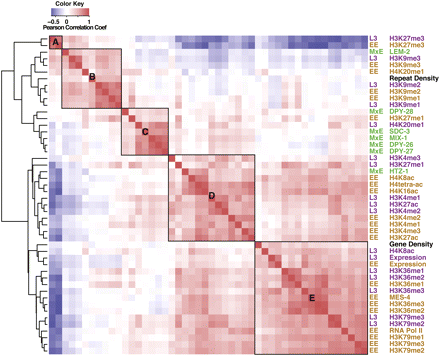
The distributions of histone marks, chromatin proteins, and genome features cluster into distinct groups. Data for all factors were clustered based on the pairwise Pearson correlations of the median signals in 1-kb windows along all chromosomes. (Red) Positive correlations; (blue) negative correlations. Factors are (brown) early embryo (EE), (green) mixed embryo (MxE), (purple) L3 larvae (L3), and (black) common features. Hierarchical clustering grouped all factors into two major groups, one related to transcriptionally repressed states (upper left) and the other related to transcriptionally active states (bottom right). The factors associated with repressed states clustered into three subgroups—A, B, and C. The factors associated with transcriptionally active states clustered into two subgroups—D associated with promoters and 5′ regions of genes and E associated with gene bodies. The dendrogram to the left of the heatmap shows the clustering tree. Pairs of horizontal branches linking two subclusters represent the distance of the two farthest separated factors in the subclusters calculated by the ChIP genome profile dissimilarity.











