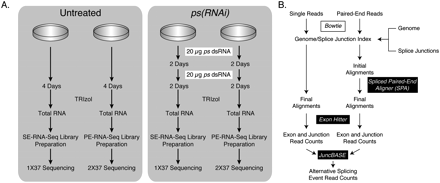
Experimental and analytical approach. (A) S2-DRSC cells were either untreated or treated with two 20-μg doses of dsRNA. After a total of 4 d of incubation with the dsRNA, total RNA was isolated and used for preparing either single-end or paired-end RNA-seq libraries. The single-end libraries were sequenced using 37–45 cycles, while the paired-end libraries were sequenced using 37 cycles for each read. The single-end sequences were trimmed from the 3′ end to a total length of 37 nt prior to alignment. (B) Sequence analysis involved aligning all reads to a combined database of the genome and splice junctions using Bowtie (Langmead et al. 2009). The paired-end alignments were further analyzed using Spliced Paired-End Aligner (SPA) to identify mate pairs that map to the same chromosome, oriented toward one another and within 200 kb of one another. The aligned reads were then analyzed using exonhitter.pl (McManus et al. 2010) to count the number of reads that mapped to exons, splice junctions, and exon–intron boundaries. The read counts were then further analyzed using juncBASE to identify alternative splicing events that were significantly different between the Untreated and ps(RNAi) samples.











