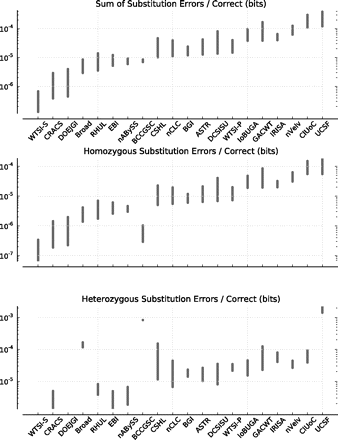
Substitution (base) errors for the top assembly from each team. (Top) Substitution errors per correct bit within all valid columns; (middle) substitution errors per correct bit within homozygous columns only; (bottom) substitution errors per correct bit within heterozygous columns only. Assemblies are sorted from left to right in ascending order by the sum of substitutions per correct bit. In each faceted plot, each assembly is shown as an interval, giving the upper and lower bounds on the numbers of substitution errors (see main text).











