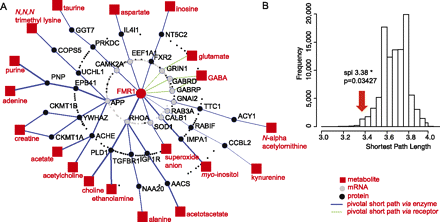
Interactome-mapping of FMR1-deficiency metabotype. (A) FMR1 integrated metabolome and interactome mapping (iMIM) network. The metabolites significantly affected in the different models (Table 1) were mapped onto the interactome, using FMRP mRNA targets and protein interactors, KEGG metabolic pathways, and neurotransmitter/receptor databases. The resulting network allows connecting of the causal gene FMR1 to the downstream metabolic consequences of its deficiency. Pivotal shortest paths via enzymes and receptors are represented in blue and green, respectively. (B) Statistical validation of the FMR1 knowledge network under the null hypothesis (H0). The average shortest path length (spl) between FMR1 and the biomarker metabolites (n = 25) was computed and compared to the distribution of average spl obtained after 100,000 H0 network simulations (i.e., networks obtained after 100,000 random selections of 25 metabolites from the entire metabolic network). This simulation under the null hypothesis shows that FMR1 appears significantly connected to the candidate biomarker metabolites, with an average distance of 3.38 hops (n = 100,000 random simulations, P = 0.03427).











