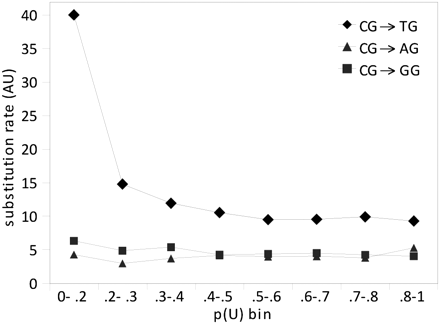
CG decay rates in orthologous human, chimp, and orang SUMIs. Substitution rates were calculated for all different types of substitutions involving the C in a CG dinucleotide ([diamonds] CG→TG, [squares] CG→AG, [triangles] CG→GG) in orthologous human, chimp, and orang SUMIs that have consistent methylation levels in the three species. Only the CG to TG substitution rate is expected to be affected by germline methylation. SUMIs in each p(U) bin have neutrophil methylation probabilities within the indicated bin boundaries in human, chimp, and orang (i.e., SUMIs considered in this analysis did not change neutrophil methylation state during these species’ evolution). Rates of CG to TG, but not to AG or GG, substitution vary as a function of methylation, in a manner consistent with neutrophil methylation states reflecting germline methylation states.











