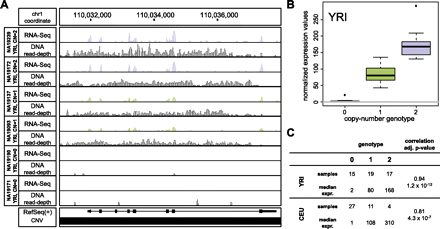
Association of a deletion with gene expression. (A) A biallelic deletion (chr1:110,024,361–110,046,935) fully deleting the GSTM1 gene was found to be significantly associated with GSTM1 transcript level in the YRI and CEU samples. RNA-seq tracks display gene expression values following normalization. DNA read-depth tracks were generated based on population-scale sequencing reads (1000 Genomes Project Consortium 2010). (B) Correlation between copy-number genotype and normalized gene expression values for GSTM1, computed based on the YRI samples. (C) Summary of observed sample abundance and median normalized expression values for each copy-number state in the YRI and CEU samples.









