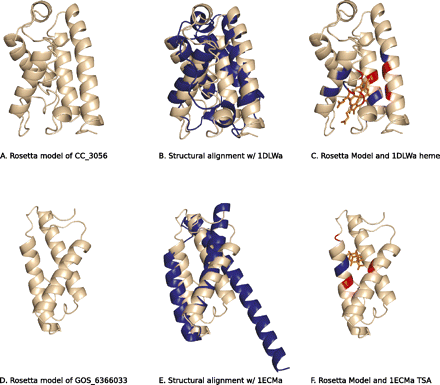
C. crescentus hypothetical protein CC_3056 is a truncated hemoglobin fold, and marine metagenome hypothetical protein GOS_6366033 is predicted to be a chorismate mutase. (A) Rosetta model of CC_3056. (B) Structural alignment of Rosetta model and 1DLWa, a truncated hemoglobin with MAMMOTH alignment Z-score of 14.39. (C) Rosetta model viewed with superimposed 1DLW heme ligand. (D) Rosetta model of GOS_6366033. (E) Structure alignment of Rosetta model and 1ECMa, a chorismate mutase with structural alignment Z-score of 9.71. (F) Rosetta model viewed with 1ECM transition state analog of chorismate. Ligand-contacting residues that are identical (blue) and similar (red) in the matched PDB structures are shown in C and F.











