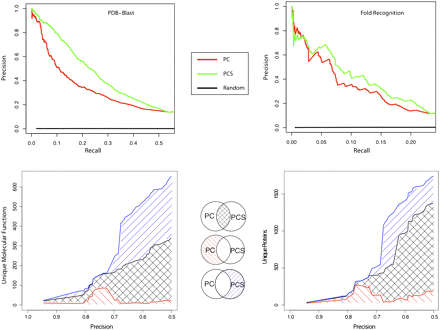
Function prediction method with structure information outperforms method without structure information. (Upper panel) Precision versus recall for eukaryotic function predictions separated by the type of structure evidence. The graph shows the precision of function predictions versus recall (fraction of all true annotations) for sequences that were structurally classified by PDB-BLAST (left) and FFAS03 (right). (Red lines) The prediction method using GO process and component (PC). (Green lines) The prediction method using GO process, component and structure (PCS). The graph shows adding structure information from PDB-BLAST and FFAS03 improves precision for function prediction. (Lower panel) Function prediction methods using structure information produce novel function predictions for a unique set of proteins. Functions were predicted for FFAS03 domains with SCOP classifications from a random sampling of 5000 proteins with known molecular function. (Black lines) Functions (lower left) and proteins (lower right) predicted by both predictor methods. (Red lines) Functions (or proteins) predicted only by PC. (Blue lines) Functions (or proteins) predicted only by PCS. The predictions made with the integration of structure with process and localization terms annotate a unique range of molecular functions and proteins that would be otherwise unreachable.











