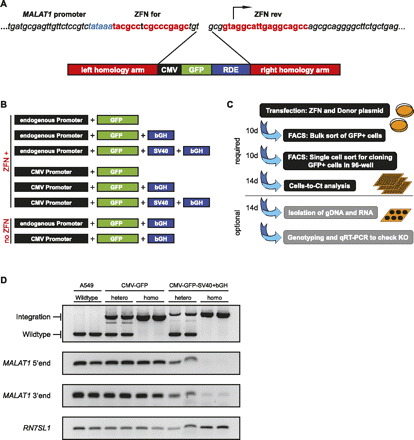
Silencing of endogenous genes by integration of RNA destabilizing elements. (A) Genomic MALAT1 region targeted by the ZFNs. The ZFNs cut between the MALAT1 TATA box (blue) and the transcriptional start site (arrow). The binding site for each ZFN is depicted in red. The site-specific integration via HR is mediated through the left and right homology arms surrounding the integration cassette containing GFP and the RDE. (B) Integration constructs used in our MALAT1 gene silencing approach. The constructs were either transfected with or without ZFNs. (C) Overview of the silencing approach. Cells were transfected with a pair of ZFNs and a repair template containing the RDE of choice, e.g., poly(A) signal. The successful gene silencing was validated via qRT–PCR and followed by genotyping. This protocol allowed the creation of single-cell clones with validated genotype and reduction in target gene expression within 6 to 8 wk. (D) Genotype–phenotype relationship of selected clones. Single cell clones (A549 wild-type, CMV-GFP, CMV-GFP-SV40+bGH) were genotyped via integration-sensitive PCR to discriminate between heterozygous and homozygous clones. The effective integration gave rise to a longer PCR product and was not present in wild-type cells. MALAT1 RNA expression was analyzed via RT–PCR using two independent primer pairs, detecting the 5′-end or the 3′-end of MALAT1. Only the homozygous, biallelic integration, including the RDE, yielded an efficient silencing of full-length MALAT1. The detection of RN7SL1 is shown as loading control.











