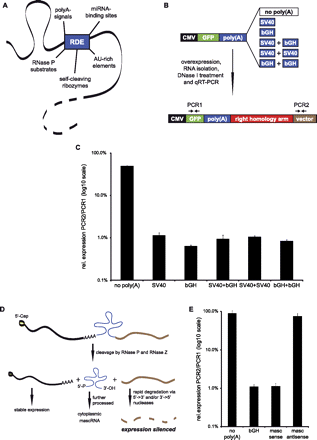
Use of RNA destabilizing elements for gene silencing. (A) RDEs that could be used for gene silencing show different silencing mechanisms. AU-rich elements and miRNA-binding sites influence the stability of the whole transcript, whereas poly(A) signals only silence downstream sequences. RNase P substrates and self-cleaving ribozymes can destabilize both upstream and/or downstream sequences, depending on the position and sequences used. (B) Different poly(A) signals were tested for their silencing potency in an in vitro combinatorial approach. A549 cells were transfected with plasmids containing combinations of poly(A) signals. Expression of GFP mRNA and vector-derived RNA was determined via qRT–PCR: Low vector encoded RNA expression indicated a strong silencing efficiency. (C) The poly(A) signals differ in silencing efficiency. The best inhibition of expression downstream from the RDE was observed with the bGH signal (mean of three experiments + SD). (D) The MALAT1-derived mascRNA sequence as RDE. Placing the mascRNA element immediately downstream from the ORF of a protein-coding gene leads to RNase P cleavage. The resulting mRNA upstream of the cleavage site is stabilized by a 5′-m7Cap and a short poly(A)-like moiety contributed by the fragment. The remaining 3′-sequence either gets degraded or is directly processed by RNase Z to yield a pre-mascRNA. The resulting 3′-end of the transcript is very unstable due to the lack of a 5′-m7Cap and a 3′-poly(A) tail, and is rapidly degraded. (E) The mascRNA element was tested for its silencing potency in the same in vitro approach used for the poly(A) signals. The 242-bp fragment was inserted immediately after the GFP ORF in sense or antisense orientation. The mascRNA element in sense orientation silenced downstream sequences as efficiently as a bGH signal (mean of three experiments + SD).











