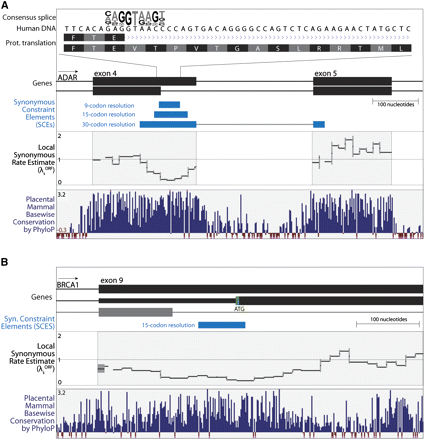
Examples of candidate synonymous constraint elements (SCEs) with likely roles in splicing and translation regulation. (A) Predicted SCEs (light blue) overlapping two isoforms of ADAR exon 4 (black) arising from an alternative splice donor site encoded within the longer exon variant. With increasing resolution, the SCE is more precisely localized to the region of overlap with the alternative splice site (motif logo for human donor sites rendered by WebLogo) (Crooks et al. 2004). The localization of the synonymous constraint to the splice site is also seen in the local synonymous rate estimate λsORF (relative to the ORF average). Note that the significant reduction in the synonymous rate is not obvious from the nucleotide-level conservation measure (dark blue, bottom panel). The extent of the predicted SCE may suggest the presence of additional splicing regulatory elements downstream from the alternative splice site. (B) Predicted SCE (light blue) overlapping an alternate translation initiation site (green) in BRCA1 encoded within exon 9 of a longer isoform. Synonymous constraint ranges from shortly upstream to immediately downstream of the alternate start codon, suggesting this region may be involved in regulating translation initiation at the alternate site. The region just upstream of the predicted SCE also shows a reduced synonymous rate (black curve) overlapping an alternative splice donor site for a third BRCA1 isoform (gray), although this reduction is not statistically significant and the third isoform is weakly supported. Annotation visualizations in Figures 3 and 4 are based on the UCSC Genome Browser (Kent et al. 2002).











