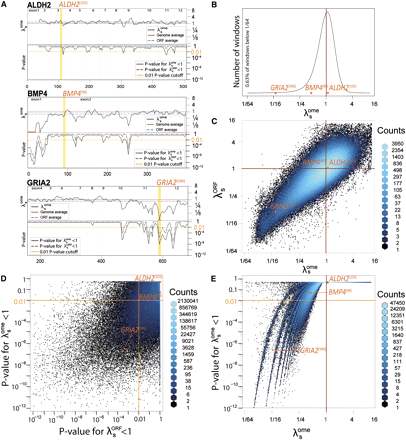
Identifying individual windows with statistically significant synonymous constraint. (A) Estimated synonymous rate relative to genome average (λsome) and corresponding P-value for the hypothesis λsome < 1 evaluated in nine-codon windows along the entire protein-coding regions of ALDH2, BMP4, and GRIA2, highlighting the windows corresponding to the three examples in Figure 1. For each plot, the top portion shows the λsome estimate for each window (black curve), the genome average (red line at λs = 1), and the ORF average (blue dashed line). The bottom portion shows the statistical significance of the reduction in the synonymous rate estimate in each window, accounting for evidence in the cross-species alignments, using a likelihood ratio test for the hypothesis λsome < 1 (continuous black curve, using the genome average as the null model), and for the hypothesis λsORF < 1 (dashed black curve, using the ORF average as the null model). (Vertical gray lines) Exon boundaries; (orange) regions where λsome drops below 1/16th toward the 5′ end of BMP4 and the 3′ end of GRIA2. (B) Overall distribution of λsome estimates for all nine-codon windows across all CCDS genes. Heavy left tail indicates an excess of windows with very low estimated synonymous rates, shifting the mean (λsome = 1) to the left of the distribution mode, which likely represents neutral rates. (C) Comparison of synonymous rates estimated relative to genome-wide (λsome) and ORF-specific (λsORF) null models, each point denoting one nine-codon window, and density of overlapping points denoted by color. Joint distribution shows that low λsome estimates also usually correspond to low λsORF estimates, and therefore that the heavy tail observed in B does not reflect regional or ORF-wide deceleration, but instead localized constraints in small windows within each ORF, also visible in the three examples of A. (D) Comparison of P-values for synonymous rate reduction with respect to genome-wide (y-axis) and ORF-specific (x-axis) null models. Candidate synonymous constraint windows are selected when synonymous rate reductions are significant at P < 0.01 with respect to both null models (orange lines). Note that many windows are significant with respect to one null model but not the other. (E) Correspondence between λsome and the associated significance estimate for the each nine-codon window. The visible stripes in this plot arise from windows that are perfectly conserved except for one, two, three, or more synonymous substitutions observed in the extant species, while the position along each stripe reflects variation in the λsome estimate and its significance, determined by the species coverage, codon composition, and observed codon substitutions in each window. (B–E) The three example regions highlighted in A are shown in each distribution and density plot, with horizontal and vertical axes aligned. The orange line in plots A, D, and E denotes the statistical significance cutoff of P < 0.01, and the red line in plots A, B, C, and E denotes the genome-wide average λsome = 1 and λsORF = 1 for B. The ALDH2[103] synonymous rate is not significantly reduced either relative to the genome or to the ALDH2 ORF; BMP4[88] is reduced relative to the genome but not relative to its ORF, which shows an overall reduced rate; GRIA2[586] is >80% reduced relative to both the genome and its ORF, resulting in significant P-values for both.











