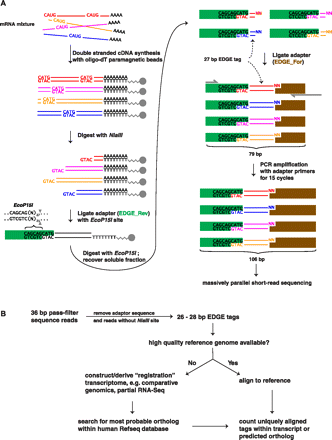
Outline of EDGE methodology and informatic pipeline for tag identification. (A) Double-stranded cDNA synthesis is performed using paramagnetic oligo(dT) beads to capture polyadenylated RNA. Next, each cDNA molecule is “anchored” by NlaIII restriction cleavage that exposes the 3′-most “CATG” site within the transcript. Following this, the EDGE_Rev adaptor (green) carrying an EcoP15I recognition site (5′-CAGCAG-3′) is ligated, and the resulting molecule is “tagged” by EcoP15I restriction digest, generating a 27-bp sequence tag. The sticky end is ligated to the EDGE_For adaptor (blue). Finally, a 15-cycle PCR amplification using adaptor-specific primers (red half-arrows) is performed to add on the additional sequence required to complete the EDGE_For adaptor and to enrich the desired final product. (B) Thirty-six base-pair pass-filter reads from the Illumina Genome Analyzer were processed to obtain EDGE tags. If a high-quality reference transcriptome was available, e.g., mouse, EDGE tags were mapped to transcript sequence and uniquely aligned tags were counted for each gene. Otherwise, EDGE tags, e.g., cheetah, were mapped to a de novo assembled reference transcriptome, e.g., cat, which acts as a scaffold to identify the orthologous gene in the organism in which the EDGE tags were derived.











