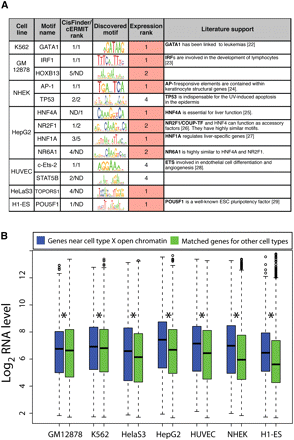
Distal cell-type selective open chromatin contains functionally relevant motifs and is linked to cell-type specific expression. (A) Top motifs enriched (P-value < 1 × 10−9) in cell-type selective open chromatin. Expression rank reflects the transcription factor's expression level in that cell type relative to all other cell types. (B) Distribution of expression values for genes closest to distal cell-type selective open chromatin sites (>2 kb from a TSS) from each cell type (x-axis) for that cell type (blue box plots). Similar distributions were calculated for these genes in the six other cell types lacking the distal open chromatin sites (green box plots). Asterisk indicates significant difference (pairwise T-tests).











