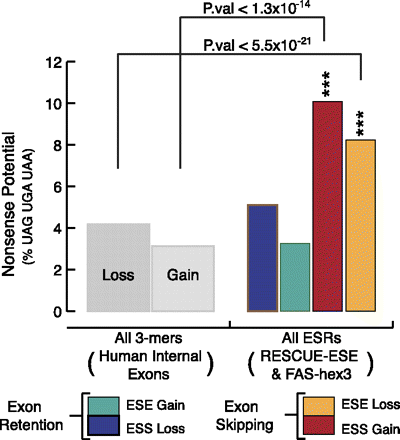
An overview of the nonsense codon sequence bias in exonic splicing regulators. Bars correspond to the nonsense-coding potential of ESR loss or gain, the proportion (expressed as a percentage) of 3-mers matching UAG, UGA, or UAA out of total 3-mers. For ESR loss, this was calculated via simulated mutation based on HGMD transition/transversion probabilities (Supplemental Fig. 2). For all human internal exonic 3-mers, the nonsense-coding potential was calculated using the same algorithm as the ESRs, except using a set of all human internal exonic sequences instead of ESR hexamers. The frequencies were normalized, and the values for the data given for ESR loss or gain were analyzed statistically (P-values from χ2 goodness-of-fit test) using an α-value of (***) 0.001.











