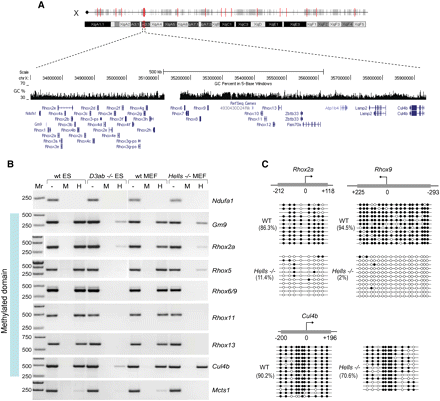
DNA methylation of Rhox gene cluster is LSH-dependent. (A) Schematic representation of the Rhox gene cluster and neighboring genes on mouse chromosome X A.33. The light-colored genes are provisional or predicted RefSeq. The GC content of the region is shown as a histogram. (B) DNA methylation at the promoters of the Rhox genes and genes neighboring the Rhox cluster was examined by PCR following digestion with methylation-sensitive HpaII (H) or methylation insensitive MspI (M) restriction enzymes of genomic DNA from wild-type and Dnmt3a/3b−/− ES cells as well as wild-type and Hells−/− MEFs. “−” represents undigested DNA. “Mr” is a DNA size marker. (C) Bisulfite DNA sequencing confirms lack of DNA methylation at Rhox2a and Rhox9 promoters, but not at the Culin4b promoter in Hells−/− compared with the wild-type MEFs. Methylated CpGs are shown as black circles. The average percentage of methylated CpGs relative to the total number of CpGs investigated by bisulfite DNA sequencing for any given sequence is shown in parentheses.











