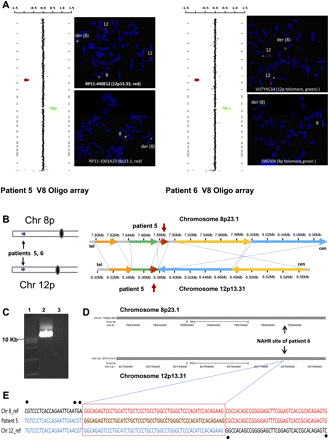
Identification of the LCR pairs acting as potential substrates for interchromosomal NAHR resulting in the t(8;12) formation. (A) CMA profiles of DNA from patients 5 (left) and 6 (right) tested on Version 8 OLIGO array (left) revealed a 7.9-Mb deletion of chromosome bands 8p23.1-pter and an 8.2-Mb duplication of chromosome bands 12p13.31-pter. The results of FISH analysis (right) with the probe RP11-440E12 (patient 5; red) or VIJTYAC14 (patient 6; green) specific for 12p33.33 indicate the presence of chromosome 12 material on the der(8), whereas the results of FISH analysis with probe RP11-1001A23 (patient 5; red) or D8S504 (patient 6; green) specific for the chromosome region 8p23 show the deletion on chromosome 8. (B) Two t(8;12) cases mapped this to NAHR substrate pair. Summary of the sequence similarity BLAST2 analysis of an ∼200-kb sequence surrounding the 8p23.1 and 12p13.31 breakpoint regions (bottom). (C) Ethidium bromide stained agarose gel image of the patient 5–specific ∼12-kb t(8;12) junction fragment amplified by long-range PCR with primers harboring trans-morphisms specific for each 8p23.1 and 12p13.31 LCR (lane 2). (Lane 1) The DNA marker with the 10-kb band indicated to the left. (Lane 3) A negative control. (D) The NAHR crossover site for patient 5 is located in a 7.7-kb subunit with 95.2% DNA sequence identity. UCSC Genome Browser view of the homologous LCR blocks in the 8p23.1 (top) and 12p13.31 (bottom) chromosome regions. (E) DNA sequence alignment of the PCR amplified translocation junction fragment for patient 6 (middle sequence). The NAHR site was narrowed to a 55-bp segment (red rectangle) with 100% sequence identity between chromosomes 8 (top, red) and 12 (bottom, blue).











