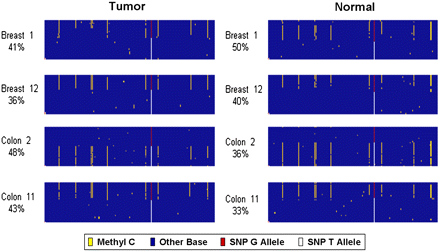
Methylation at the H19 imprinted locus. Data from four patients who were germline heterozygous for a SNP (rs2251375) in this locus. The sequencing reads are aligned as rows in each panel. Each base in the read is color-coded to indicate the sequence: (yellow) a methylated cytosine; (blue) all other bases. The position of the SNP is indicated by the red and white column: (red base) reads from the G allele; (white base) reads from the T allele. The percentage of reads for each patient that are from the G allele is listed below the patient identifier for each sample. As expected for an imprinted locus, methylation is observed on one allele in both the tumor (left panels) and the adjacent normal tissue (right panels) for each patient. Both alleles and both methylated and unmethylated molecules were amplified and sequenced efficiently from this locus in all samples.











