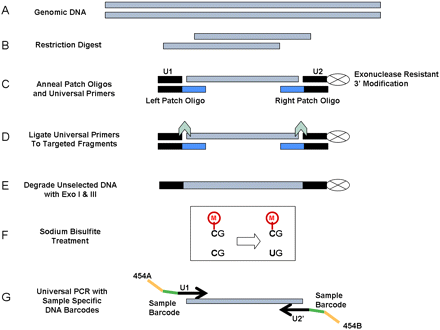
Bisulfite Patch PCR. (A,B) Genomic DNA restriction digest. (C) Anneal patch oligos and universal primers specifically to the ends of desired fragments. (D) Ligate universal primers (U1 and U2) to targeted fragments. (E) Degrade unselected DNA with exonucleases. Targeted loci are protected from exonuclease by 3′ modification on U2. (F) Treat with sodium bisulfite to convert unmethylated cytosine to uracil, leaving methylated cytosine intact. (G) PCR all loci simultaneously with universal primers tailed with sample-specific DNA barcodes and sequencing machine primers (454A and 454B). Pool PCR products from all samples together for sequencing.











