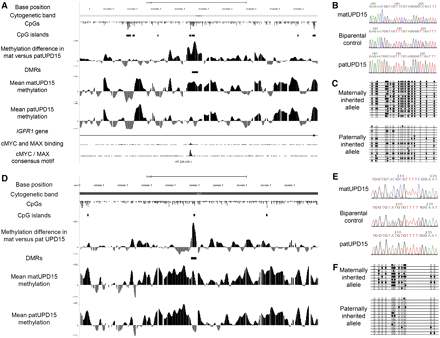
Identification of differentially methylated regions within intron 2 of the IGF1R gene in 15q26.3 and distal to GABRG3 in 15q12. (A,D) Images show the smoothed mean log2 ratios from three individuals with paternal UPD15, three individuals with maternal UPD15, the difference between these two profiles, and putative DMRs identified by statistical analysis uploaded as custom tracks in the UCSC Genome Browser. The DMR within intron 2 of the IGF1R gene contains the sequence RACCACGTGGTY, corresponding to the methylation-sensitive consensus binding motif for MYC/MAX (Solomon et al. 1993), and in vivo binding of both MYC and MAX to this site in K562 cells is confirmed by chromatin immunoprecipitation with massively parallel sequencing (ChIP-seq). Each plot shows a 50-kb window centered on each DMR: (A) chr15:97,203,000–97,253,000; (D) chr15:25,575,000–25,625,000. Also shown are the genomic coordinates (hg18), cytogenetic band, CpG dinucleotides, and CpG islands defined using epigenetic criteria (Bock et al. 2007). (B,E) Sequencing after bisulfite modification of the DNA, which converts unmethylated cytosine residues to uracil, confirms that these two loci both show differential methylation between matUPD15 and patUPD15 samples. A biparental control shows the expected pattern with an approximately equivalent mix of both methylated and unmethylated alleles. (C,F) Individual bisulfite-treated alleles from HapMap individual NA12753 were isolated by cloning and divided by parental origin using heterozygous informative SNPs within each amplicon. In each case, the maternally derived allele is predominantly methylated, while the paternally derived allele is predominantly unmethylated, in agreement with both the array data and bisulfite sequencing of UPD15 cases. Each line represents a separate clone. (●) Methylated CpGs; (○) unmethylated CpGs.











