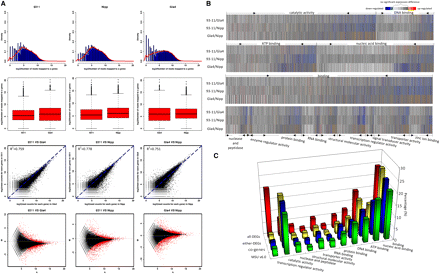
Analyses of DEGs between indica and japonica. (A) The top row of graphs show log2 ratios of the expressed genes in 93-11, Nipponbare, and Guangluai 4, respectively. The second row of box-and-whisker plots show log2 transformed values of the expressed genes in the three rice varieties. The third row of graphs show the scatter plot comparing the gene expression levels between 93-11 and Gla4, between 93-11 and Nipp, and between Gla4 and Nipp, respectively. The bottom row of graphs show the genes identified as differentially expressed by Fisher's exact test (red dots). (B) Cluster display of 34,738 coexpressed genes by GO analysis. Differentially expressed genes are shown in red (up-regulated) and blue (down-regulated). Genes with similar expression levels are shown in gray. (C) Genes are categorized on the basis of expression support. (MSU) MSU annotated ; (co-genes) coexpressed genes; (either-DEG) differentially expressed genes in either two varieties; (all-DEG) differentially expressed genes in all three varieties. The percentage of each molecular function for the gene categories is shown.











