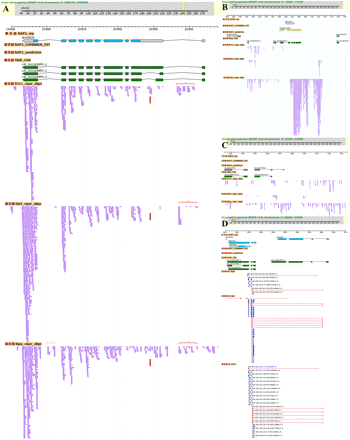
Examples of identified novel AS patterns and nTARs. (A) Novel exons (brackets) and alternative exons (vertical arrows) in the LOC_Os12g38850 transcript among 93-11, Guangluai 4, and Nipponbare are shown. Exons (filled boxes and horizontal arrows) and introns (lines) predicted by gene models (RAP2 and TIGR) were indicated. The RNA-seq short reads were indicated by purple lines. (B) Strong transcriptional activity was detected by the RNA-seq reads (purple lines) in the Nipponbare nearby gene LOC_Os12g01290. (C) nTARs identified by RNA-seq reads (purple lines) in Nipponbare and 93-11 as compared with the gene prediction models. (D) Novel exon–exon splicing junctions were identified in the three varieties.











