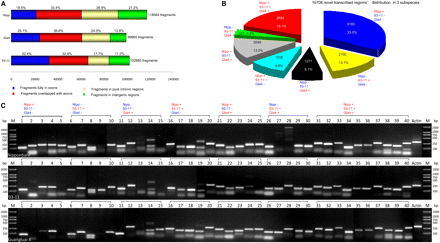
Characterization and classification of nTARs. (A) Distribution of continuous transcribed fragments according to the MSU annotated gene models. (B) The 15,708 nTARs were classified in seven categories based on the positive (+) or negative (−) detection of the transcribed fragments in three rice varieties, 93-11, Guangluai 4, and Nipponbare. (C) RT-PCR validation of 40 randomly selected transcripts from the seven categories (indicated at top) of nTARs was carried out using the total RNAs of the three rice varieties, Nipponbare, 93-11, and Guangluai 4, as indicated. Amplification of the actin fragment in RT-PCR was used as control.











