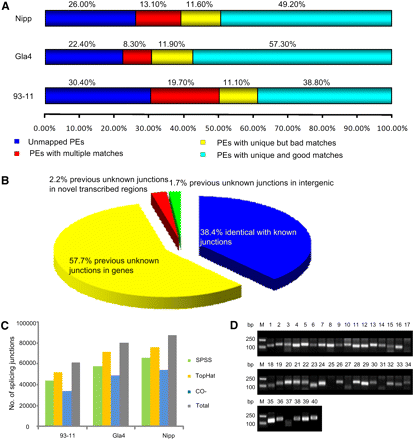
Summary of RNA-seq mapping data. (A) Overall mapping results of paired-end reads (PEs) referring to the Nipponbare genome sequence. (B) Classification of the identified exon–exon junctions based on the known splicing junctions of the MSU annotated gene models. (C) Splicing junctions identified by SPSS and TopHat. (D) RT-PCR validation of 40 randomly selected novel identified exon–exon junctions were carried out by using the total RNAs of Nipponbare. Amplification of the actin fragment in RT-PCR was used as control.











