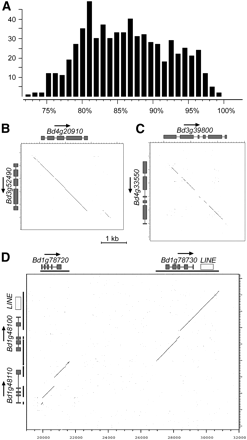
(A) Levels of DNA sequence identities between 534 gene copies from the donor region and acceptor sites where DNA sequence conservation allowed a reliable alignment. (X-axis) Level of DNA sequence identity; (y-axis) the number of gene pairs in that identity range. (B) DotPlot alignments of donor region and acceptor site. In recent duplications, the gene and its flanking regions are well conserved, and the borders of the duplicated region can clearly be identified. (Gray boxes) Exons of the predicted gene. (C) With time, duplicated regions diverge and sequence conservation is basically limited to the protein-coding exons. (D) DotPlot alignment of a duplicated fragment containing two genes. The duplicated fragments are indicated as black bars underneath the genes. The interruptions in the conserved parts are caused by insertions or deletions that occurred after the duplication. Molecular dating of the conserved LINE (white box) indicates that the duplication occurred ∼1.3 Mya.











