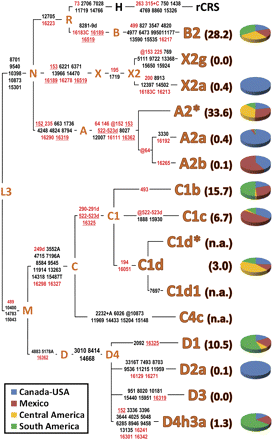
MtDNA tree encompassing the roots of all known Native American haplogroups. The distinguishing mutational motifs for the 15 known Native American haplogroups are reported on the branches. The position of the revised Cambridge reference sequence (rCRS) (Andrews et al. 1999) is indicated for reading off-sequence motifs. Mutations in the control region are in red, while mutations in the coding region are listed in black; they are transitions unless a base is explicitly indicated. The prefix @ designates reversions, while suffixes indicate transversions (to A, G, C, or T), indels (+, d). Recurrent mutations within the tree are underlined. The percent frequency of each Native American haplogroup in the entire double continent is reported in parentheses and has been obtained from the Sorenson Molecular Genealogy Foundation Mitochondrial (SMGF) mtDNA database (http://www.smgf.org) (entire control region) excluding all non-Native American mtDNAs. For each haplogroup, the relative frequencies in northern America (Canada and USA), Mexico, Central America, and South America are reported in different colors in the corresponding pie chart. Some haplogroups are completely absent in the SMGF mtDNA database because either they are extremely rare (X2g) or harbor a restricted distribution range (D3 in the Eskimos and Aleuts). For C4c, C1d*, and C1d1 frequency values are not available (n.a.) due to the lack of distinguishing control region mutations, but the overall C1d incidence (C1d* plus C1d1) is reported.











