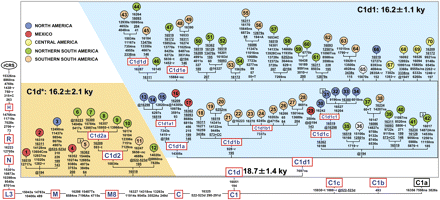
Detailed tree of C1d in the context of haplogroup C1. All 73 C1d mtDNA sequences (63 novel and 10 published) are complete except for samples 36 and 65, for which only coding-region data are available. The basal motifs for Native American haplogroups C1b and C1c are also included together with the motif of the Asian-specific branch C1a. The position of the revised Cambridge reference sequence (rCRS) (Andrews et al. 1999) is indicated for reading off-sequence motifs. Mutations are shown on the branches; they are transitions unless a base is explicitly indicated. The prefix @ designates reversions, while suffixes indicate transversions (to A, G, C, or T), indels (+, d), gene locus (∼t, tRNA; ∼r, rRNA; ∼nc, noncoding region outside of the control-region), synonymous or nonsynonymous changes (s or ns), and T/C heteroplasmy (Y). Recurrent mutations within the phylogeny are underlined. We have followed the guidelines for standardization of the alignment in long C stretches (Bandelt and Parson 2008), but disregarded any length variation in the C-stretch between nucleotides 303 and 315, with the exception of the well-known 315+C insertion. Additional information regarding each mtDNA is available in Supplemental Table S1. Coalescence times shown for C1d, C1d*, and C1d1 are maximum-likelihood (ML) estimates, while the corresponding averaged distance (ρ) accompanied by a heuristic estimate of SE (σ) are shown in Table 1. As for the geographic affiliation (top left corner), North America refers to USA and Canada; northern South America refers to Colombia, Venezuela, Ecuador, Peru, and Brazil; southern South America corresponds to Chile, Argentina, Uruguay, and Paraguay.











