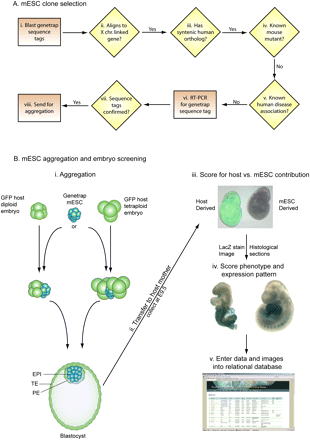
An overview of the screening method. (A) mESC GT clone selection involved blasting of sequence tags to transcripts from the X chromosome (i). All BLAST (Altschul et al. 1990) hits >95% identical were then compared with the human X chromosome (ii). Those genes with a syntenic homolog were then filtered for any known mouse mutants and known links to human disease (iv,v). GT lines were ordered and quality controlled by RT-PCR to confirm the sequence tag prior to aggregation (vii,viii). (B) Two different aggregation techniques (i) were used with similar results and efficiency (Table 1). mESCs were aggregated with either one diploid or two tetraploid host embryos and cultured overnight. After overnight culture, the colony of mESC preferentially forms the epiblast of the embryo and displaces the host cells to the primitive endoderm and trophectoderm of the blastocyst. mESCs contribute only to the epiblast (EPI) that will generate the embryo proper, while the host (all tetraploid and a large proportion of diploid) embryos generate the primitive endoderm (PE) and the trophectoderm (TE). Aggregated embryos were transferred to pseudopregnant mothers and collected at E9.5 (ii). Dissected embryos were scored for mESC contribution by observing GFP fluorescence (iii). The embryo on the left is GFP positive and has diploid host embryo contribution, while the embryo on the right is GFP negative and is fully mESC-derived (iii). Fully mESC-derived embryos were fixed and stained for beta-galactosidase (lacZ) activity. After staining, the embryos were digitally imaged and scored for expression and phenotype (iv). All digital images and aggregation data were entered into a relational database (v).











