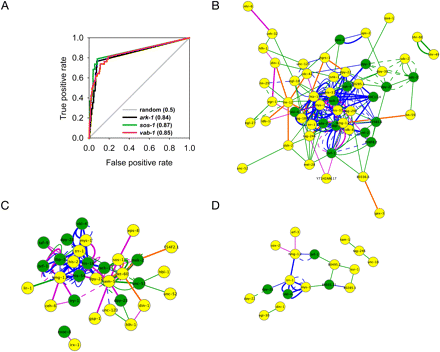
Predicting and validating novel genetic modifiers for three signaling genes in C. elegans. New candidate genetic modifiers were predicted for mutations in three signal transduction genes by ranking all genes in the genome by their functional coupling to the known genetic modifiers for each gene as shown schematically in Figure 2A. The top ranked approximately 90 genes that had not been previously tested for their ability to interact were then tested. (A) Cross-validated ROC curves for each of the three genes' previously known genetic modifiers. AUCs are indicated after each gene name in key. (B–D) Previously known (yellow nodes) and newly verified (green nodes) genetic modifiers of mutations in vab-1 (B), sos-1 (C), and ark-1 (D) are plotted as networks, showing the high network connectivity among the genes that led to their prediction as genetic modifiers. Edges show WormNet 2 predicted functional couplings between the modifier loci, derived from evidence in C. elegans (green edges), D. melanogaster (orange edges), S. cerevisiae (blue edges), and H. sapiens (purple edges). Physical interaction evidence is shown as thick lines, cocitation evidence as dashed lines, and all other evidence as solid lines. Gene networks were plotted using Cytoscape 2.5.2. (Cline et al. 2007).











