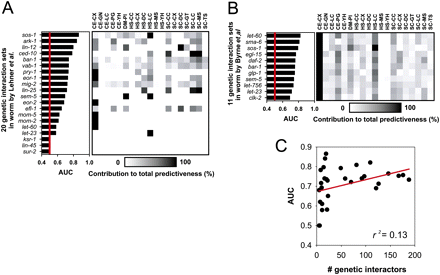
High predictive power for genetic modifiers stems from a wide variety of data types integrated into WormNet. Both independent screens for genetic modifier for worm orthologs of human disease genes by Lehner et al. (2006a) (A) and by Byrne et al. (2007) (B) shows that groups or genetic modifiers (labeled by sharing genetic interaction partner disease gene names at y-axis) with high AUC scores (random expectation is indicated by red line where AUC = 0.5) are supported by contribution (degree of contribution is measured and indicated as in Fig. 1D) of diverse data types (listed at x-axis with same code scheme as Fig. 1A). (C) Predictability does not depend on seed set size, seen by a low correlation between the number of known genetic modifiers (number of seed genes) and predictability (indicated by AUC).











