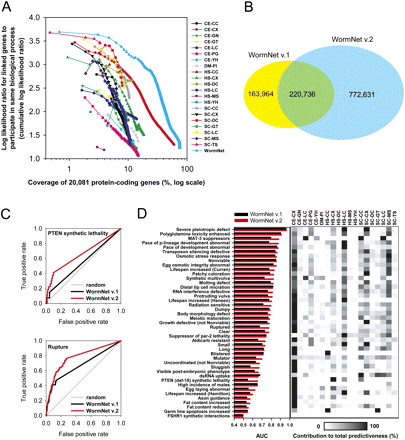
Improved performance of the updated gene network for C. elegans, WormNet version 2. (A) Benchmarking results of network by each 21 data sets and one by their integration (WormNet). Each data set is indicated by XX-YY code, in which XX represents origin species of data—CE, C. elegans; DM, Drosophila melanogaster, HS, Homo sapiens, SC, Saccharomyces cerevisiae—and YY represents data type—CC, cocitation; CX, coexpression; GN, gene neighbors; GT, genetic interaction; LC, literature curated protein–protein interaction; PG, phylogenetic profiling; YH, high-throughput yeast two-hybrid interaction; PI, protein interaction; DC, domain co-occurrence; MS, mass spectrometry analysis; TS, inferred interaction from protein tertiary structure. The x-axis represents coverage of 20,081 protein-coding worm genes by different components of WormNet version 2 (log scaled); the y-axis represents predictive performance of the network and components, measured as the cumulative log likelihood for linked genes to participate in same Gene Ontology biological process. (B) Venn diagram between old and new WormNet linkages. The new WormNet more than doubles the number of linkages, with >57% (220,736 of 384,700) of the WormNet v.1 linkages recapitulated in v.2. (C) Improved predictability for RNAi phenotypes by WormNet v.2 is illustrated by ROC curve analysis for two RNAi phenotypes. A version of WormNet omitting literature-based genetic interaction datasets (CE-CC and CE-GT) was used for all ROC analyses in Figures 1–4, to minimize the possibility of circular reasoning when predicting published genetic interactions. (D) The improved predictability of WormNet v.2 (red bars) over v.1 (black bars) is illustrated by a comparison of AUC scores for 43 different RNAi phenotypes. For each RNAi phenotype (rows), the supporting power of each data set (columns, labeled as in A) to the prediction was measured as a fractional score—the sum of log likelihood scores of all supporting evidence for that data set divided by the sum of log likelihood scores across all data sets. The degree of contribution is indicated by grayscale, where the higher support the darker indicator. Various data sets including those from other species make significant contribution for predictability for worm RNAi phenotypes.











