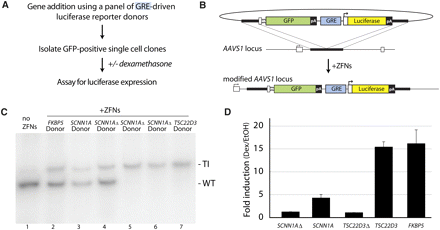
Use of ZFNs to generate a panel of U2OS cells carrying glucocorticoid receptor reporter constructs at AAVS1, and their functional characterization. (A) Outline of experiment. (B) Schematic of donor design and of the AAVS1 locus following GFP marker/GRE luciferase reporter addition. Gene elements are represented in the same way as in Figure 1B. (C) Genotype at the AAVS1 locus for clones carrying reporters with GREs derived from genes indicated above each lane. The position of the wild-type (WT) and transgenic (TI) chromatid is indicated to the right of the gel. The “SCNN1AΔ” and “ TSC22D3Δ ” donors have the GR binding site deleted by site directed mutagenesis. Note that the transgenic chromatid is amplified less efficiently than the wild-type one due to a difference in size (see Supplemental Discussion). (D) The single-cell-derived clones genotyped in C were treated with vehicle (EtOH) or dexamethasone (dex); induction of the luciferase reporter was measured and is shown as fold activation by dex over treated with vehicle only.











