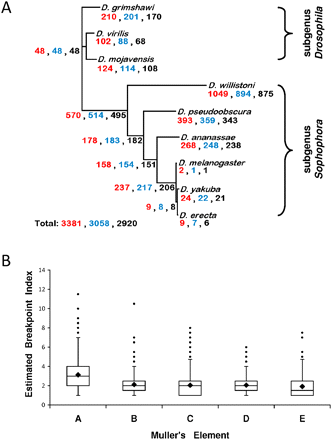
Gene order evolution in the genus Drosophila. (A) Magnitude of independent gene anchor (IGA) order evolution and its phylogenetic distribution across the genus Drosophila. The magnitude of change is expressed as the number of inversions, as estimated under maximum parsimony using multiple genome rearrangement (MGR) (Bourque and Pevzner 2002), for three different synteny definitions: (red) conserved gene order and orientation (GOO); (blue) conserved gene order (GO); and (black) only overall local contiguity (OLC). (B) Box plot of the average breakpoint index (BI) across Muller's elements as inferred from MGR. (Box) Interquartile range (from the 25th to the 75th percentile, i.e., 50% of the values); (line across the box) median; (rhombus) mean; (whiskers) maximum and minimum values with the exception of the outliers (solid circles). The BI equals the number of times each of the edges of a HCB or singleton has been involved in a chromosomal rearrangement since the ancestral genome to the genomes of the species studied here divided by 2. Mann-Whitney U tests were performed between pairs of Muller's elements, and the resulting P-values were adjusted with the Bonferroni correction. All the comparisons involving Muller's element A were statistically significant (P < 1.0 × 10−4); the only additional pairwise comparison that entails statistically significant differences involves Muller's elements B and E (P = 4.5 × 10−3). The results shown correspond to the GO synteny definition.











