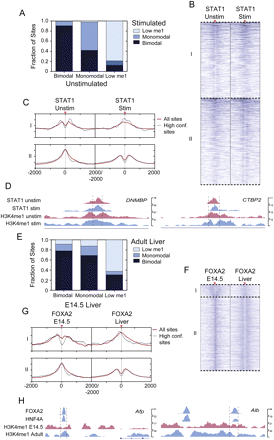
H3K4me1-marked nucleosome occupancy regulates transcription factor bound cis-regulatory site function. Analysis of H3K4me1 transitions for transcription factor occupied loci during (A–D) an IFNG-stimulated signaling event, or (E–H) liver development. (A) STAT1 site class transitions in the IFNG-stimulated and unstimulated shared peaks, or (E) FOXA2 adult occupied loci in E14.5 hepatocytes and adult liver. Categories I and II in the heatmaps in B and F represent bimodal–monomodal and monomodal–bimodal transitions for STAT1 and FOXA2 sites, respectively. Average H3K4me1 enrichment profiles of category I or II sites are plotted in red, with dotted lines showing the average profiles for high confidence peaks. UCSC mm8 Genome Browser screenshots of representative loci that are bound by (D) STAT1 in the IFNG-stimulated and unstimulated cells that transition from bimodal to monomodal or vice versa, or (H) by FOXA2 and HNF4A in adult liver but transition from bimodal to monomodal or vice versa. The dotted lines in D and H mark 1-kb regions.











