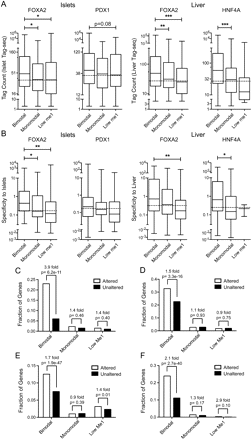
Bimodal sites regulate target gene expression. Box-whisker plots of islet or liver (A) Tag-seq counts or (B) specificity against 202 other mouse SAGE libraries, for expressed genes associated with a bimodal, monomodal, or low H3K4me1 site in the FOXA2 (islet or liver), PDX1, or HNF4A peak sets. Expressed genes have a tag count >5 in the particular Tag-seq library. Statistically significant differences were detected using a Kruskall-Wallis non-parametric test with a Dunn's multiple comparison correction, with a P < 0.05 threshold. (*) P < 0.05; (**) P < 0.01; (***) P < 0.001. Fractions of RefSeq genes with bimodal, monomodal, or low H3K4me1sites whose expression was significantly altered or not in (C) PancChip 6.1 (http://www.betacell.org/ma/) comparisons of Foxa2 deficient (Foxa2loxP/loxP, Pdx1-CreERT2, tamoxifen treated) versus control (Foxa2loxP/loxP, tamoxifen treated) islets (Gao et al. 2007). (D) Tag-seq analysis of islets treated with siRNA's targeting Pdx1 versus with siRNA's targeting Ppib. (E) Agilent 4×44k array comparisons of Foxa2 deficient (Foxa2loxP/loxP, Alfp-Cre) versus wild-type livers (Bochkis et al. 2009). (F) Agilent Whole Mouse Genome Microarray comparisons of Hnf4a deficient (Hnf4aloxP/loxP, Alb-Cre) versus wild-type livers (Holloway et al. 2008). P-values were determined using Fisher's exact test.











