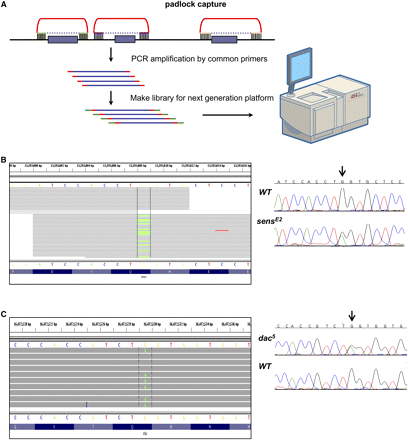
Two known mutations were detected using padlock capture sequencing. (A) Flowchart of mutation detection by padlock capture and next-generation sequencing technologies. Padlock capture technology requires probes that contain two target-specific capturing arms (green) connected by a common linker (red). The unique targeting arms of individual targeting oligonucleotides are designed to hybridize immediately upstream and downstream from each exon (purple bars) of interest. Hybridization to genomic DNA is followed by an enzymatic gap filling and ligation step, such that a copy of the sequence of interest is incorporated into a circle (purple dashed line). Then the enriched DNA is PCR amplified and is used to prepare libraries for the next-generation platform sequencing. (B) sensE2 and (C) dac5 mutation detection. Reads alignment at the mutation is shown between the vertical lines. Each read is drawn as a gray line, and bases that are different from the reference are colored. Sanger sequencing is conducted to confirm the mutation with the heterozygous mutated base indicated by the arrow.











