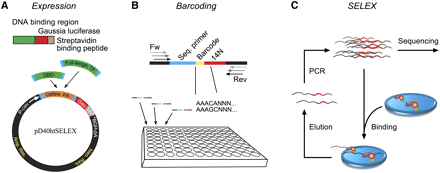
Schematic description of the high-throughput SELEX process. (A) Protein expression. (Top) Proteins are expressed as fusion proteins with SBP-tagged Gaussia-luciferase. (Bottom) The GATEWAY recombination cloning system is used to transfer DNA sequences encoding DBDs or TFs from donor-vectors to the pD40htSELEX expression vector. (B) Ligand design that accommodates multiplexing of samples using barcodes. Each DNA ligand contains a 14-bp randomized region (14N), and a 5-bp barcode (Barcode) that uniquely identifies the individual SELEX sample. To increase specificity, each barcode differs from all other barcodes by at least 2 bp. These variable sequences are flanked by constant sequences that include an Illumina Genome Analyzer sequencing primer site (Seq. primer) and bridge amplification primer binding regions (Fw, Rev; arrows), which are extended in their 5′ regions to accommodate partially nested primers (used in successive SELEX rounds). (C) Basic principle of high-throughput SELEX. A double-stranded DNA mixture containing all possible 14-bp sequences (from B) is incubated with a DNA-binding protein immobilized into a well of a 96-well plate, resulting in binding of DNA to the protein. After washing and elution, the resulting population of more specific sequences is amplified by PCR and subjected to high-throughput single-molecule sequencing. The specificity of the TF can then be constructed by iterating the process and calculating the abundance of distinct sequences after different numbers of cycles. In each cycle, multiple reactions are mixed into a single sequencing lane, and the TFs are identified using the barcode sequences.











