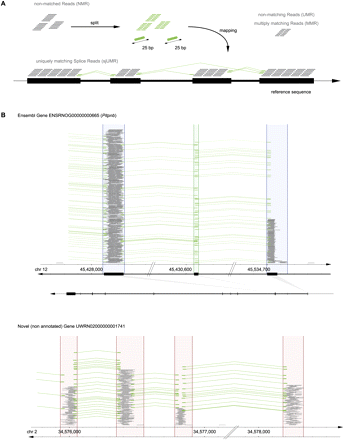
Agnostic splice junction (SJ) mapping. (A) Agnostic, i.e., de novo genome-wide SJ discovery from split reads (25 bp + 25 bp). NMR were split into 25-mer halves, which were matched independently to the genome reference sequence. Reads whose halves each matched uniquely to the genome defined SJs de novo; these were termed uniquely mapping splice junction reads (sjUMR). (B) SJ validation on known genes and their application to define connections of novel exons and exon borders. (Green) SJ-spanning uniquely mapping reads (sjUMR; mapped as described in the Methods section). Most SJs were supported by multiple sjUMR (28.6 on average), as depicted here by piles of green lines. (Top panel) A known gene (Pitpnb ENSRNOG00000000665) is shown here, demonstrating how sjUMR mapped precisely to the border of exons as annotated in the reference genome. (Bottom panel) sjUMR connecting novel UMR clusters to groups, thereby defining new genes. In some instances, UMR cluster borders fall into introns (as a result of the inclusiveness of the sliding-window algorithm), as depicted here (right sides of second and third cluster). In those cases, exon borders were defined precisely by sjUMR. Note (top panel, second exon) that very short (<50 bp) exons that were undetectable by UMR could be defined through mapping of SJ reads.











